By Heidi Toth
NAU Communications
Ecologists frequently tag animals in the wild, monitoring their movements and activities to gain a greater understanding of the species’ behaviors and lifestyle.
Benjamin Koch is one of the first ecologists to propose doing the same with bacteria.
Koch, an assistant research professor in the Center for Ecosystem Science and Society (Ecoss) and the Department of Biological Sciences, led a team of biologists and ecologists to study antibiotic-resistant bacteria through an ecological lens. They outlined new ways to use genomic “tags” to track the bacteria. More and more scientists are paying attention to these bacteria, which are increasing in prevalence as antibiotic use in humans and food animals, or animals intended for food consumption, spikes throughout the world.
It’s a complicated situation, which is why Koch and his colleagues, including Ecoss director Bruce Hungate, wanted a different take on it.
“We clearly need to use antibiotics more responsibly,” Koch said. “However, limiting the spread of antibiotic resistance also requires knowing the extent to which antibiotic-resistant microbes move among farms, the environment and people.”
Koch’s study, which will be published in the August issue of Frontiers in Ecology and the Environment, applied ecological principles to newly acquired genomic data on antibiotic-resistant bacteria. He and his colleagues looked at existing data from scientific literature as well as data on antibiotic use and resistance collected by the U.S. Department of Agriculture and the Centers for Disease Control and Prevention.
Armed with these data, they applied concepts usually reserved for more traditional ecological issues, such as how non-native species invade an ecosystem, to identify the conditions that enable antibiotic-resistant bacteria to successfully colonize in humans.
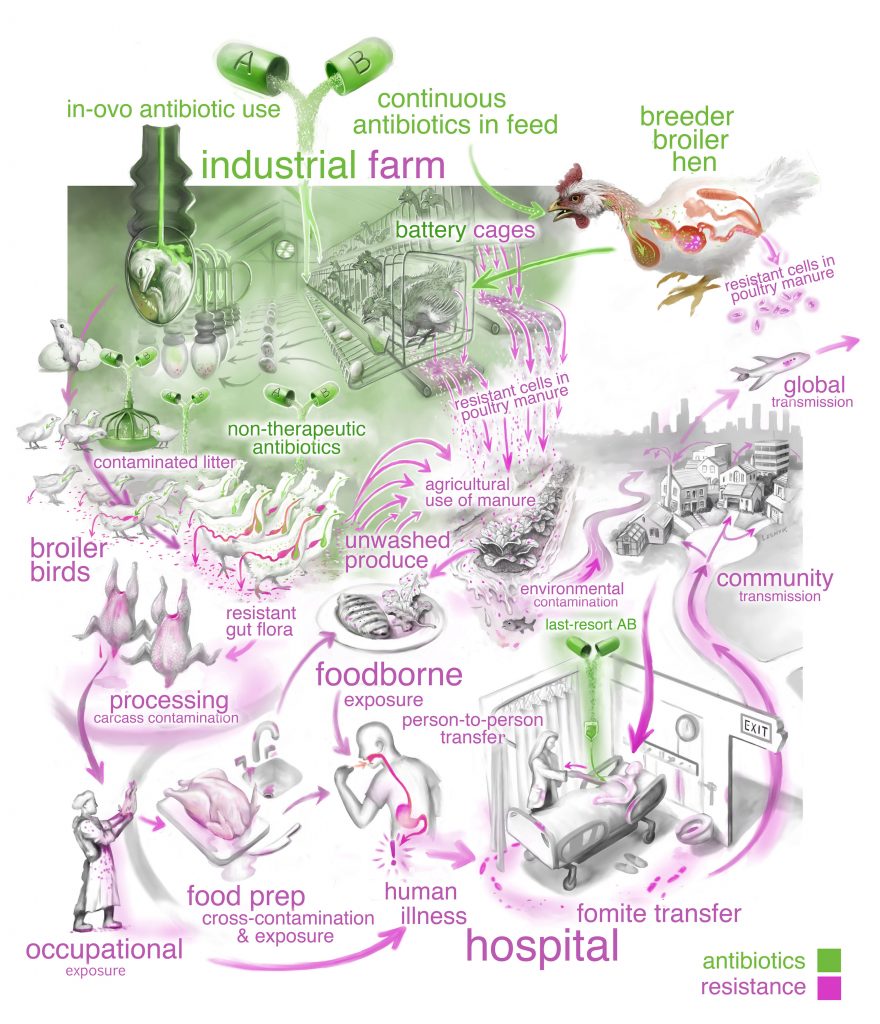
The data they gathered highlight the pathways antibiotic-resistant bacteria use to move between food animals and humans. By applying basic ecological principles—essentially looking at the data with the perspective of how species move, reproduce, survive and interact—Koch and his colleagues can help organize what scientists know to better monitor, understand and manage the spread of antibiotic-resistant bacteria.
“This highly detailed view into the transmission patterns and lifestyles of antibiotic-resistant bacteria is revealing new insights for controlling their spread,” Koch said. “The approach of combining ecological theory with genomic data could lead to new strategies for controlling antibiotic resistance.”
The next step of this research will be to get a “microbe’s eye view” of the environmental factors and biological interactions of the bacteria, which will include studying samples of resistant bacteria throughout the food-production chain. They also plan to look at person-to-person transmission of other types of antibiotic-resistant pathogens.
This study and the studies that follow are aimed to help scientists find new ways of combating antibiotic resistance beyond just developing more and stronger antibiotics. Those alternative approaches could include implementing more efficient hygienic measures that target specific antibiotic-resistant strains or strengthening the human microbiome so it is better able to keep the antibiotic-resistant bacteria in check.
“Melding ecological theory with increasingly prevalent genomic data on antibiotic resistance has the potential to revolutionize how we monitor and manage antibiotic resistance,” Koch said. “It may also give us powerful new tools for slowing the spread of antibiotic resistance.”
Koch and Hungate also collaborated with Lance Price, director of the Center for Microbiomics and Human Health at the Translational Genomics Research Institute in Flagstaff and a professor of environmental and occupational health at George Washington University.